Note
Go to the end to download the full example code
2.23 Kuramoto-Sivashinsky - Compiled methods
This example implements a scalar PDE using a custom class with a numba-compiled method for accelerated calculations. We here consider the Kuramoto–Sivashinsky equation, which for instance describes the dynamics of flame fronts:
\[\partial_t u = -\frac12 |\nabla u|^2 - \nabla^2 u - \nabla^4 u\]
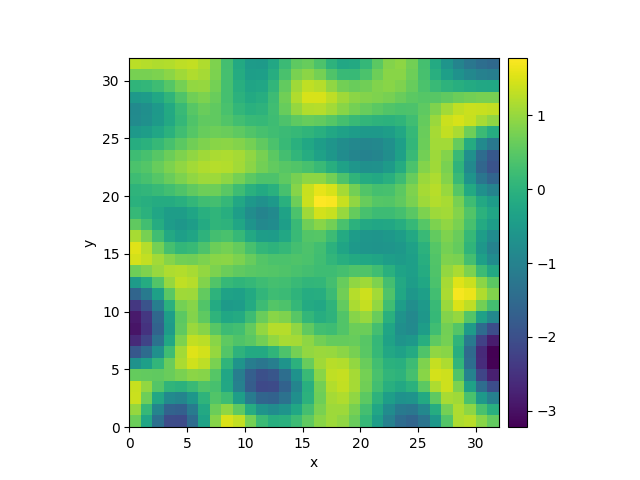
0%| | 0/10.0 [00:00<?, ?it/s]
Initializing: 0%| | 0/10.0 [00:00<?, ?it/s]
0%| | 0/10.0 [00:10<?, ?it/s]
0%| | 0.01/10.0 [00:11<3:09:55, 1140.67s/it]
0%| | 0.02/10.0 [00:11<1:34:52, 570.35s/it]
2%|▏ | 0.18/10.0 [00:11<10:22, 63.38s/it]
76%|███████▌ | 7.62/10.0 [00:11<00:03, 1.50s/it]
76%|███████▌ | 7.62/10.0 [00:11<00:03, 1.50s/it]
100%|██████████| 10.0/10.0 [00:11<00:00, 1.14s/it]
100%|██████████| 10.0/10.0 [00:11<00:00, 1.14s/it]
import numba as nb
from pde import PDEBase, ScalarField, UnitGrid
class KuramotoSivashinskyPDE(PDEBase):
"""Implementation of the normalized Kuramoto–Sivashinsky equation"""
def __init__(self, bc="auto_periodic_neumann"):
super().__init__()
self.bc = bc
def evolution_rate(self, state, t=0):
"""implement the python version of the evolution equation"""
state_lap = state.laplace(bc=self.bc)
state_lap2 = state_lap.laplace(bc=self.bc)
state_grad_sq = state.gradient_squared(bc=self.bc)
return -state_grad_sq / 2 - state_lap - state_lap2
def _make_pde_rhs_numba(self, state):
"""nunmba-compiled implementation of the PDE"""
gradient_squared = state.grid.make_operator("gradient_squared", bc=self.bc)
laplace = state.grid.make_operator("laplace", bc=self.bc)
@nb.njit
def pde_rhs(data, t):
return -0.5 * gradient_squared(data) - laplace(data + laplace(data))
return pde_rhs
grid = UnitGrid([32, 32]) # generate grid
state = ScalarField.random_uniform(grid) # generate initial condition
eq = KuramotoSivashinskyPDE() # define the pde
result = eq.solve(state, t_range=10, dt=0.01)
result.plot()
Total running time of the script: (0 minutes 11.575 seconds)