Note
Go to the end to download the full example code
2.25. Custom PDE class: SIR model
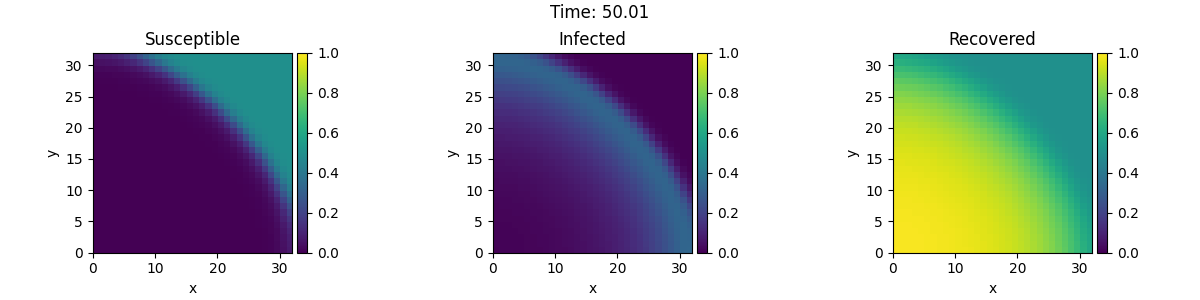
This example implements a spatially coupled SIR model with the following dynamics for the density of susceptible, infected, and recovered individuals:
\[\begin{split}\partial_t s &= D \nabla^2 s - \beta is \\
\partial_t i &= D \nabla^2 i + \beta is - \gamma i \\
\partial_t r &= D \nabla^2 r + \gamma i\end{split}\]
Here, \(D\) is the diffusivity, \(\beta\) the infection rate, and \(\gamma\) the recovery rate.
0%| | 0/50.0 [00:00<?, ?it/s]
Initializing: 0%| | 0/50.0 [00:00<?, ?it/s]
0%| | 0/50.0 [00:00<?, ?it/s]
0%| | 0.02/50.0 [00:01<45:05, 54.13s/it]
0%| | 0.04/50.0 [00:01<22:38, 27.18s/it]
1%| | 0.31/50.0 [00:01<03:02, 3.68s/it]
3%|2 | 1.48/50.0 [00:01<00:44, 1.09it/s]
8%|7 | 3.96/50.0 [00:01<00:21, 2.16it/s]
15%|#5 | 7.58/50.0 [00:02<00:14, 3.01it/s]
24%|##3 | 11.96/50.0 [00:03<00:11, 3.27it/s]
32%|###2 | 16.05/50.0 [00:04<00:09, 3.63it/s]
41%|####1 | 20.73/50.0 [00:05<00:07, 3.69it/s]
50%|##### | 25.01/50.0 [00:06<00:06, 3.89it/s]
60%|#####9 | 29.77/50.0 [00:07<00:04, 4.05it/s]
69%|######9 | 34.74/50.0 [00:08<00:03, 4.04it/s]
78%|#######8 | 39.2/50.0 [00:09<00:02, 4.15it/s]
88%|########7 | 43.99/50.0 [00:10<00:01, 4.12it/s]
97%|#########6| 48.35/50.0 [00:11<00:00, 4.20it/s]
97%|#########6| 48.35/50.0 [00:12<00:00, 3.99it/s]
100%|##########| 50.0/50.0 [00:12<00:00, 4.12it/s]
100%|##########| 50.0/50.0 [00:12<00:00, 4.12it/s]
from pde import FieldCollection, PDEBase, PlotTracker, ScalarField, UnitGrid
class SIRPDE(PDEBase):
"""SIR-model with diffusive mobility"""
def __init__(
self, beta=0.3, gamma=0.9, diffusivity=0.1, bc="auto_periodic_neumann"
):
super().__init__()
self.beta = beta # transmission rate
self.gamma = gamma # recovery rate
self.diffusivity = diffusivity # spatial mobility
self.bc = bc # boundary condition
def get_state(self, s, i):
"""generate a suitable initial state"""
norm = (s + i).data.max() # maximal density
if norm > 1:
s /= norm
i /= norm
s.label = "Susceptible"
i.label = "Infected"
# create recovered field
r = ScalarField(s.grid, data=1 - s - i, label="Recovered")
return FieldCollection([s, i, r])
def evolution_rate(self, state, t=0):
s, i, r = state
diff = self.diffusivity
ds_dt = diff * s.laplace(self.bc) - self.beta * i * s
di_dt = diff * i.laplace(self.bc) + self.beta * i * s - self.gamma * i
dr_dt = diff * r.laplace(self.bc) + self.gamma * i
return FieldCollection([ds_dt, di_dt, dr_dt])
eq = SIRPDE(beta=2, gamma=0.1)
# initialize state
grid = UnitGrid([32, 32])
s = ScalarField(grid, 1)
i = ScalarField(grid, 0)
i.data[0, 0] = 1
state = eq.get_state(s, i)
# simulate the pde
tracker = PlotTracker(interval=10, plot_args={"vmin": 0, "vmax": 1})
sol = eq.solve(state, t_range=50, dt=1e-2, tracker=["progress", tracker])
Total running time of the script: ( 0 minutes 12.563 seconds)